Understanding
Vector Technologies
What are Vectors and Why do You Need Them?
Key advantages include:

Introduction to AAV
As their name suggests, AAVs are “associated” with adenovirus as they require co-infection with an adenovirus to replicate in host cells. These viruses are also known as “helper viruses” in the context of producing recombinant AAV.
AAV2 Genome Map (Schematic)
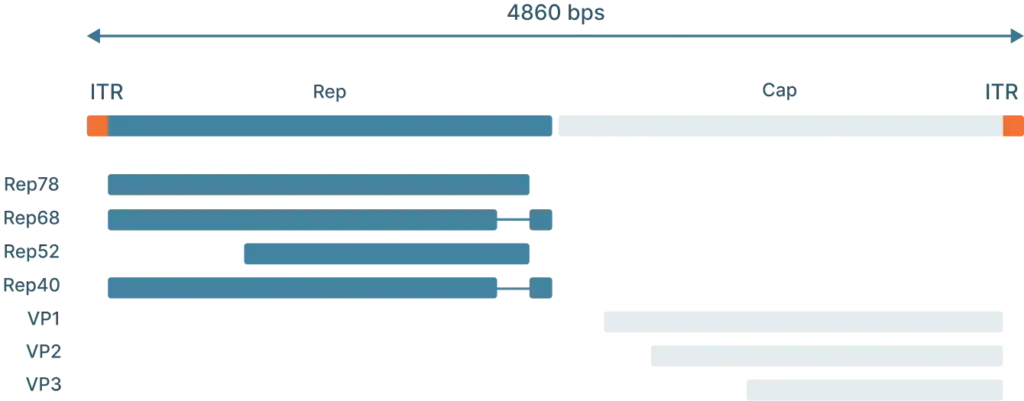
To understand how scientists use AAVs in the lab, we first need to understand the genome and life cycle.
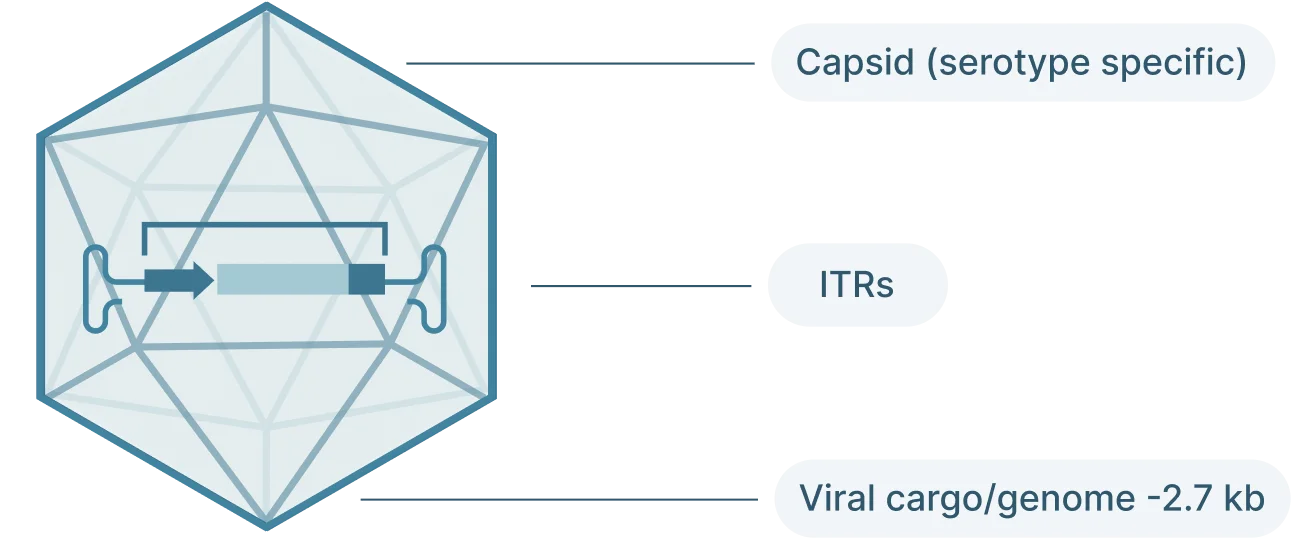
The AAV genome is viral genome is single-stranded DNA, ~ 4.7 kb long and consists of three main parts:
- Inverted terminal repeat (ITRs): Located at both ends of the DNA strand, the ITR sequences flank the viral genome.
- Rep: The Rep (replication) genes encodes 4 non-structural proteins involved in genome replication and packaging. However, Because the AAV genome doesn’t encode a polymerase, AAVs still require the host cell’s polymerase to replicate its genome.
- Cap: The Cap (capsid) genes encod 3 viral capsid proteins that form the outer “shell” of the virus encapsulating the AAV genome and determines tissue tropism by allowing the virus to bind to specific cell receptors
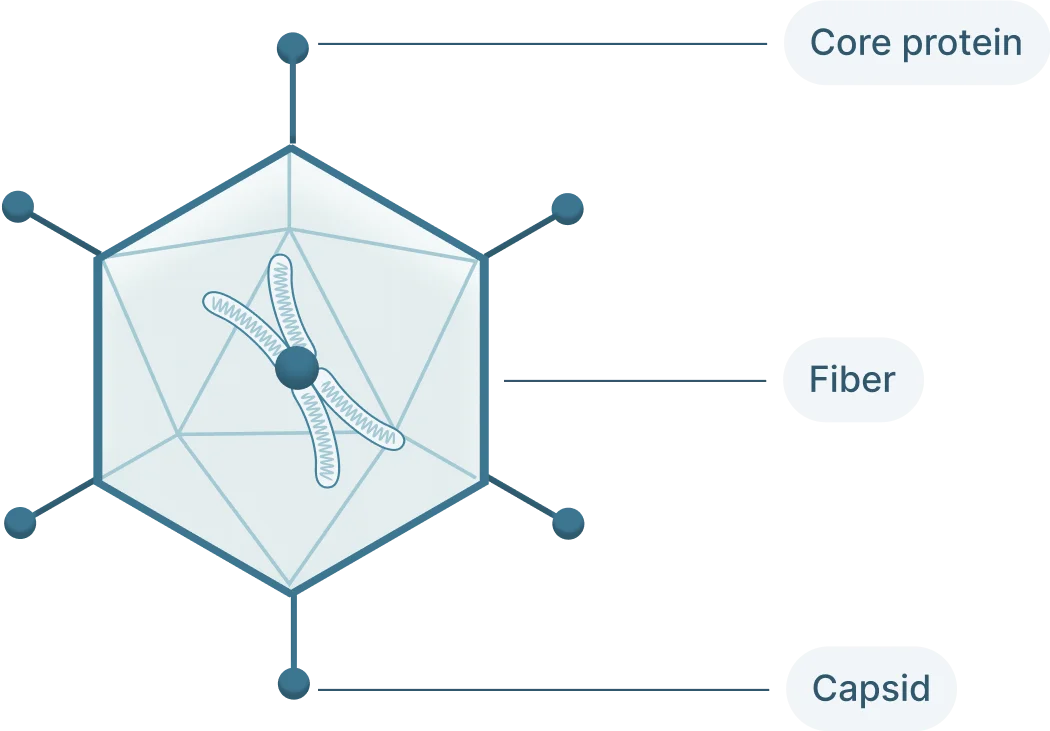
AAVs depend on their viral capsid to enter host cells. The capsid interacts with receptors on the host cell’s surface before it can be internalized into the cell. The exact receptors required depend on the AAV’s serotype, AAV2 requires heparan sulfate proteoglycan. Once internalized, the virus enters the nucleus of the cell.
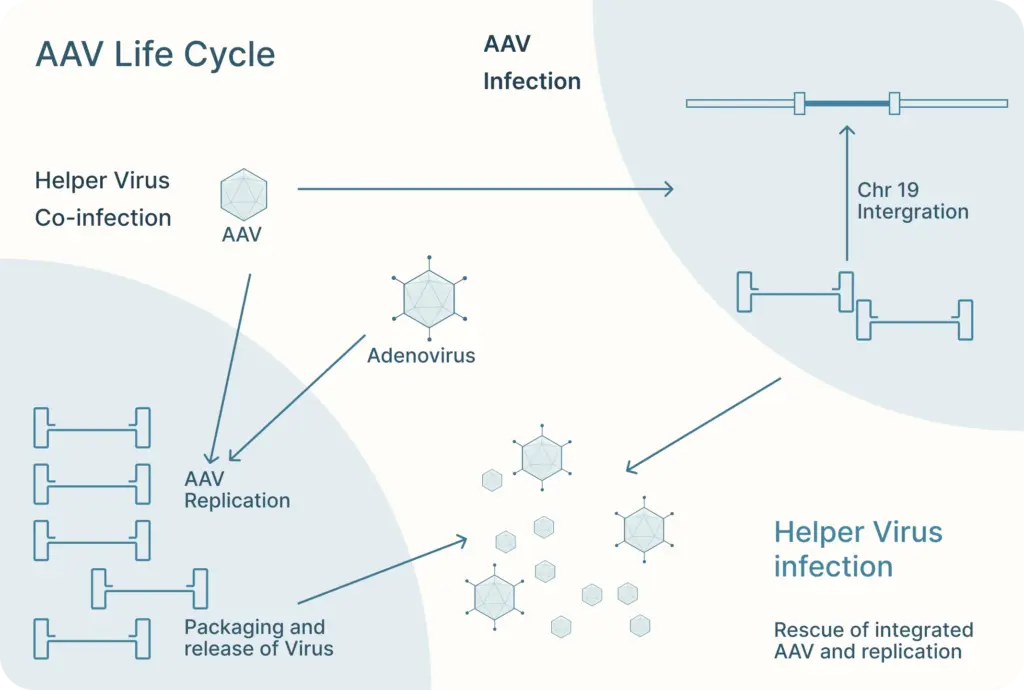
There, AAV can follow one of two distinct and interchangeable pathways of its life cycle: the lytic cycle and the lysogenic cycle.
If the AAV is infected along with a helper virus, such as adenovirus or herpes simplex virus, the AAV enters the lytic cycle. During this, the AAV gene expression program is activated and new AAV particles are produced within the cell. These cells burst and release newly formed viral particles into the environment where they can infect other cells to continue the lytic cycle.
The lysogenic life cycle occurs when the AAV enters the host cell without co-infection of a helper virus. Its gene expression program is auto-repressed and the AAV genome integrates into a specific region on the long arm of human chromosome 19. This integration involves the AAV ITRs and Rep proteins (Rep78, Rep68, Rep52 and Rep40). The AAV can also enter the lytic cycle after being integrated into the host chromosome when a helper virus is introduced. Rep proteins can excise the AAV genome out of the host chromosome, where they can be replicated and packaged in the lytic cycle.
There are many characteristics that make AAVs a great tool for gene editing.: This includes:
- Low pathogenicity: Wild-type AAVs are non-pathogenic. For example, 80-90% of adults are seropositive with AAV2, but infection hasn’t been associated with any symptoms or disease
- Low immunogenicity: AAV produces the lowest immune response compared to other viral vectors.
- Broad tropism: AAV can infecttransduce many tissues including liver, muscle, heart, kidney, retina, lungs and brain a wide range of tissues.
- Multiple serotypes: The AAV has more than 20 different serotypes that allow for tissue- and wde-spread specificity increases specificity to certain cell types or can be more broad. AAVs can also infect both dividing and quiescent cells.
Vectors/genes needed
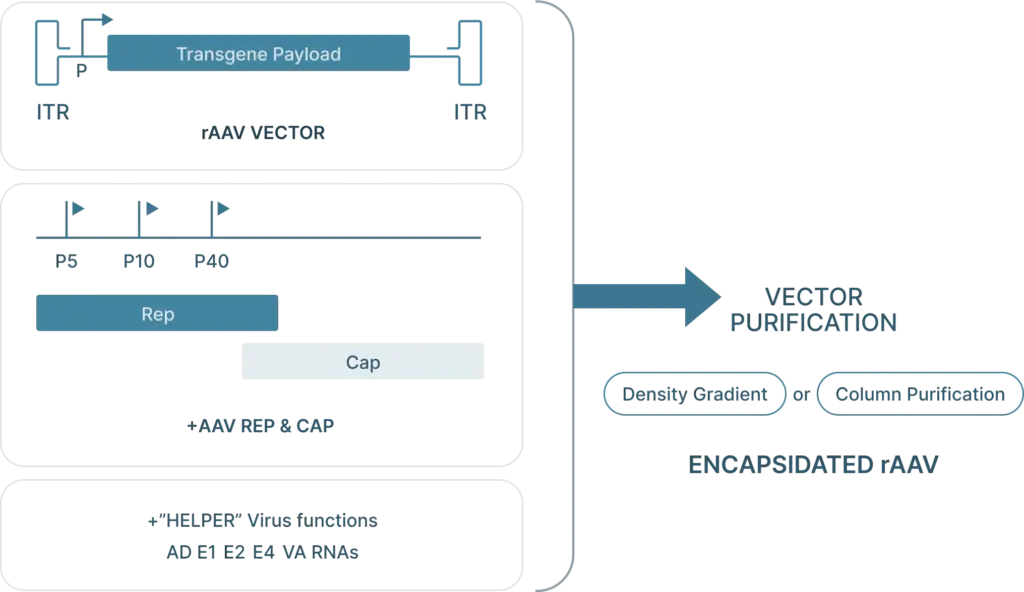
To generate recombinant AAV (rAAV), the following are required:
- Cis plasmid: The transfercis plasmid contains the two ITRs from the AAV genome, with the Rep and Cap genes replaced with the desired “payload.” This payload is the gene that you want to introduce into your cells (along with a selective or ubiquitous promoter upstream of the gene for expression) and must be less than 4.5 kb in length so that it can fit inside the capsid.
- Transfer plasmid: The transfer plasmid provides the Rep and Cap genes that direct the AAV serotype.were originally on the AAV. This plasmid does not include the e ITRs. aren’t included in this plasmid so these genes won’t be packaged into viral particles.
- Helper virus genes: There are five helper viral genes needed to create rAAVs. Three of these genes are typically provided on a helper plasmid (E4orf6, E2a, and VA RNA). Two other helper genes (E1a and E1b55k) are typically expressed by the HEK293 cell line commonly used to produce rAAV. It’s possible to use one plasmid that provides both the adenoviral helper genes as well as Rep and Cap genes.
AAV Production Protocol
The first step is to subclone the gene of interest (GOI), shRNA, or gRNA in the final pAAV cis-plasmid which is then amplified in preparation for viral production. The common approach to produce rAAV is by a triple transfection (cis-, trans- and helper- plasmids) in a producer cell line. Vector Biolabs currently uses HEK293 cells. The HEK293 cells are harvested ~3 days post-transfection followed by purification by CsCl combined with ultracentrifugation. Once the rAAV is purified Vector BioLabs performs a series of bioanalysis, for example: (1) viral purity, (2) determination of physical titer, (3) aggregation by DLS and SLS, (4) empty/full capsid ratio by UV/Vis, (5) endotoxin testing by LAL and more.
rAAV2 Vector Production
There are also engineered serotypes that have been designed to infect new tissues or evade the immune system. Many of these capsids are engineered using directed capsid evolution, capsid DNA shuffling, or peptide library insertion technologies. , E,g, AAV-DJ is a hybrid capsid created through DNA shuffling that combines eight different AAV capsids AAV-DJ, combines capsid elements from eight different serotypes and has higher transduction efficiency in vitro compared to any naturally of its parental occurring serotypecapsids.
AAV can be tailored by over 20 different serotypes (naturally occurring and engineered) that allow for (1) tissue-specific and wide-spread targeting, and (2) systemic injections that can penetrate the blood brain barrier.
The question is: among these serotypes of AAV, which serotype(s) is appropriate or best for a particular study model?
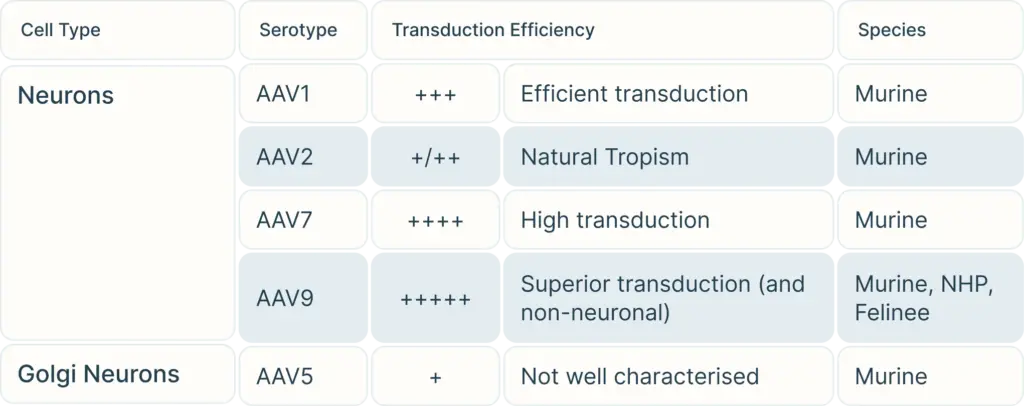
To begin, consider the cell type you are targeting. Our Vector Selection Guide can help you choose serotypes depending on your cell type and species. Once you have selected your potential serotypes, you can carry out experiments using a reporter gene such as GFP or LacZ to verify transduction efficiency in your exact study.
However, one major drawback to AAVs is that they can only hold trangenes DNA payload less than 4.5 kb in length. Larger transgene can be spliced into multiple fragments for each fragment to be carried by its own AAV virus, and then combine with a second and even third AAV virus carrying carrying all fragments of the gene for injection. Beyond that lengthAlternatively, you can also’d need to consider another viral vector such as lentivirus or adenovirus to carry a larger gene.
Introduction to Adenovirus and Recombinant Adenovirus
Adenovirus have been developed as gene delivery tools for various vaccine and gene therapy (ex: to treat cancer) applications against infectious diseases.
This article gives an overview of the wild-type adenovirus genome and life cycle, and how recombinant adenoviruses are produced and purified.
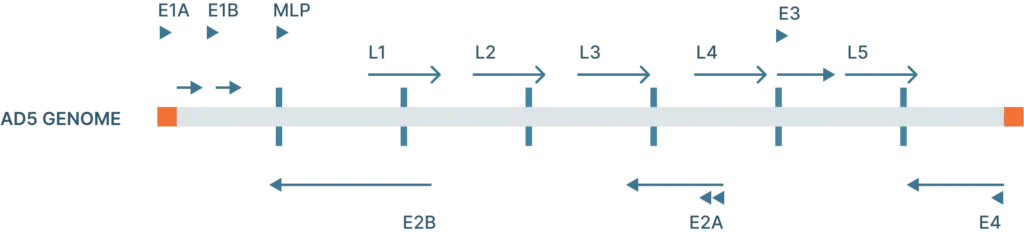
The adenovirus genome is linear double stranded of DNA that isof 26-46 kb in length and codes for 23-46 proteins (Figure 1). These genes are split into two categories:
- Early genes (E1-4): codes for proteins involved in viral transcription, viral DNA replication, and suppressing the host response
- Late genes (L1-5): codes for viral capsid components and capsid assembly
These genes are flanked by inverted terminal repeats (ITRs) that are involved in initiating viral DNA replication and are binding sites for transcription factors.
After creating the transgene construct for adenoviral vectors, they can then be produced and purified (Figure 4). The first step is to transfect the cell line (ex: HEK293 cells) with the adenoviral plasmid containing the transgene. There, the virus can infect the cells and replicate. After a few weeks, the cells are harvested and lysed to create the low-titer virus stock. Next, the virus is amplified to increase their titer by infecting more cells through multiple rounds of amplification.
Once amplified, viral particles can be purified using ultracentrifugation in CsCl or iodixanol density gradients and quantified.

The adenovirus genome is linear double stranded of DNA that isof 26-46 kb in length and codes for 23-46 proteins (Figure 1). These genes are split into two categories:
- Early genes (E1-4): codes for proteins involved in viral transcription, viral DNA replication, and suppressing the host response
- Late genes (L1-5): codes for viral capsid components and capsid assembly
These genes are flanked by inverted terminal repeats (ITRs) that are involved in initiating viral DNA replication and are binding sites for transcription factors.
Adenovirus
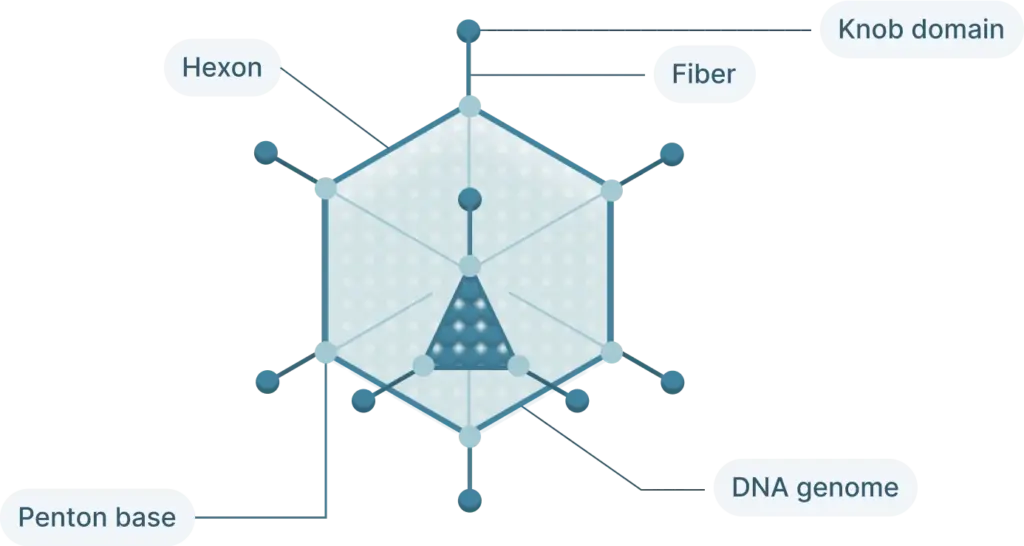
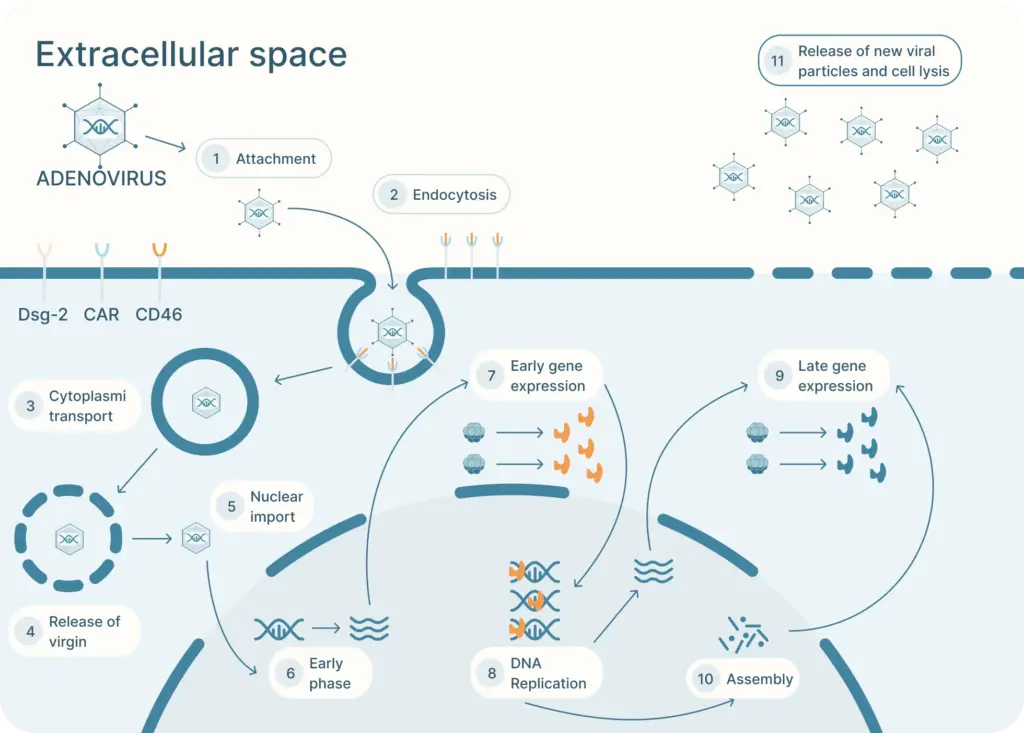
The adenovirus life cycle (Figure 2) involves five main stages, the first begins with viral entry into the host cell. The adenoviral knob domain of the capsid binds to the appropriate cell receptor The virus’s knob domain binds to the appropriate cell receptor, which depends on the adenovirus type. Most vectors are based on adenovirus type 5 (Ad5) which use the Coxsackie-Adenovirus Receptor (CAR) to enter cells.
A secondary interaction then occurs between the virus’s penton base and host integrins. Next, the virus is internalized into an endosome, which acidifies. This partially disassembles the capsid. The virus then escapes near the nucleus and releases its DNA into the nucleus via a nuclear pore complex.
Once in the nucleus, the viral genes are transcribed. The early genes are transcribed first. Then DNA replication occurs which is followed by expression of the late genes. The virus assembles and bursts from the host cell.
There are many advantages to using an adenovirus to introduce genetic material into host cells.
Recombinant adenovirus:
- Has the ability to infect most mammalian cell types (both replicative and non-replicative)
- Accommodates reasonably large transgenes (up to 8 kb for first generation adenovirus and up to 36 kb for gutless adenoviral vectors). This contrasts to the 4.7 kb capacity for AAV.
- Allows high expression of the recombinant protein
- May be grown at high titer (10E10 VP/mL, which can be concentrated up to 10E13 VP/mL)
- Is well tolerated, with post-infection viability of the host cells being almost 100%
- Does not integrate into the host chromosome and does not inactivate genes or activate oncogenes
- Induces a strong immune response, which can be beneficial for vaccination applications
However, because adenovirus surface proteins elicit a strong immune response, recombinant adenoviral vectors can only be expressed transiently (~2 weeks) before the host clears the virus from the cell. This limits the repeated use of the vector. In addition, pre-existing immunity in humans (especially towards Ad5) limits the effectiveness of adenoviral vectors. To circumvent this, it’s possible to develop adenoviral vectors based on less common serotypes.
Since scientists have begun developing recombinant adenoviral vectors, they’ve been making iterative improvements (Figure 3).
The first generation of adenovirus vectors began with deleting the E1 and E3 gene. Deleting these genes gives the adenoviral vector 8kb in packaging capacity and renders them unable to replicate on their own so that they can be safely used as a gene delivery tool. HEK293 cells, which are used to produce adenovirus, supply the E1 gene, but it’s possible that recombination between the cell’s DNA and the viral genome could create replication competent adenovirus. To circumvent this, scientists have developed 911 cells to express E1 in human embryonic retinoblasts, but these cells are not as widely available as HEK293 cells. The E3 gene, which is involved in evading host immunity, is not essential for viral production so it can be left out entirely.
Subsequently, a second generation of adenovirus vectors was developed. In addition to the removal of the E1 and E3 gene, these vectors also do not encode E2 and E4. This increases the capacity of the vector to ~14 kb.
Another more recent variation of adenoviral vectors are the gutless adenoviral vectors (also called high capacity adenoviral vectors, or HC-Adv). These vectors remove all of the viral coding genes on the transfer plasmids, leaving behind only the ITRs and the packaging domain, which aids in recognition of the viral genome during packaging. These vectors can hold up to 35 kb. Producing gutless adenoviral vectors requires a helper virus to carry all of the necessary coding regions of the virus.
